I decided to dig a little deeper into the climatic adaptation of Himalayan maize. You may remember from my last post on this that Genesys has 96 maize accessions from over 2000 masl in the Himalayas, collected at some 50-odd unique localities. When I ran these accessions through the Subsetting Tool in Genesys, I got the following histogram.
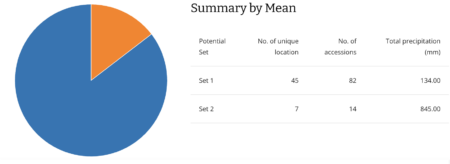
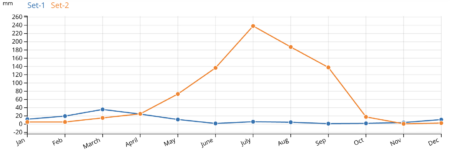
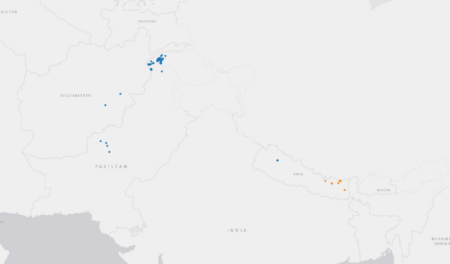
What struck me — and surprised me — was the spike of sites way at the left hand of the precipitation plot. So I took a closer look at the results of the subsetting analysis. And the clustering algorithm it uses to look for similar sites did in fact identify two climatically quite different groups of locations: 45 of the unique high altitude maize collecting sites (the blue ones) are indeed drier than the other 7 (in orange).
Much drier. (And also colder actually, but that’s another story.)
They’re the ones mainly collected in Pakistan and Afghanistan.
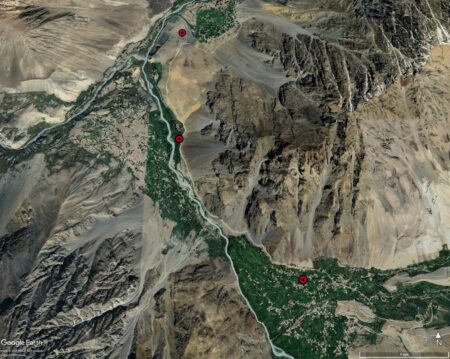
Now, I don’t know whether these areas really get 135 mm of annual precipitation, which seems really low, and in any case the agriculture there is clearly rainfed.
But those maize samples, mainly now conserved at CGN in the Netherlands incidentally, the results of something called the 1976 Netherlands-Pakistan Expedition by the Stichting voor Plantenveredeling, do seem to have some very unique adaptations.