- Seed collection and processing practices affect subsequent seed storage longevity in durum wheat and wild relatives. Immature seeds can still usefully be harvested for long-term storage of properly handled.
- Two-step drying of soya bean seed germplasm often improves subsequent storage longevity. “Proper handling” includes drying at higher temperatures.
- Seed-stored transcript integrity as a molecular indicator of viability in conserved common bean germplasm. mRNA degradation predicts loss of seed viability.
- Developing a cryopreservation protocol for the conservation of coconut palm (Cocos nucifera L.) using a novel type of explant, meristematic clumps. Who needs seeds anyway?
- Pollen cryobanking at the USDA-ARS National Laboratory for Genetic Resources Preservation. Well, who needs meristematic clumps?
- Mapping pea seed composition through strategic selection of accessions from the Nordic gene bank. Image analysis can be used to maximise diversity in nutritional composition in pea seeds, thus facilitating use of genebank collections. Can’t do that with pollen, I suspect.
- A small-scale assessment of the availability of EURISCO accessions. Facilitating use needs all the help it can get.
- Strengthening national genebanks through genomics and regional collaboration: Lessons from Latin America and the Caribbean. I guess genomics capacity could help with use.
- Enhancing farmers’ access and use of conserved germplasm for improved food security and climate resilience: The case of sorghum at Kenya’s national genebank. Genomics unavailable for comment. Farmers, on the other hand….
- Linking the ICRISAT Genebank to Poverty Reduction and Welfare in Malawi. Facilitating use by farmers is important.
- Farmers as breeders and seed producers: Insights from 30 years of scaling up seed clubs in Vietnam. It’s super cool when farmers organize. Including for genebanks.
- Elephant ear yam Xanthosoma robustum Schott (Araceae), a neglected crop native to Central America. Needs more attention from genebanks. And farmers and their clubs for that matter.
- Plant genebank of Sudan: Towards recovery from the wreckage of war to a new era of further capacity development based on lessons learnt from similar situations. We must de-risk genebanks. Wouldn’t want to lose all those elephant ear yam collections we’ll be making.
Brainfood: Diversity of Sugarcane, Rice, Lentils, Olives, Sweetpotato, Cassava, Beans, Buckwheat, Pigeon pea, Landscapes
- The genomic footprints of wild Saccharum species trace domestication, diversification, and modern breeding of sugarcane. The genome of modern sugarcane is a mosaic of wild introgressions, including one from an unknown source.
- Evolutionary histories of functional mutations during the domestication and spread of japonica rice in Asia. Selection by biotic stresses acted differently on standing variation in rice across geographic regions. Colour me surprised.
- Ancient DNA from lentils (Lens culinaris) illuminates human-plant-culture interactions in the Canary Islands. Local lentils trace back a thousand years in the Canaries.
- An olive parentage atlas: founder cultivars, regional diversification, and implications for breeding programs. Modern cultivars derive from a surprisingly small set of founding genotypes…
- Intraspecific variation and phenotypic plasticity of olive varieties in response to contrasting environmental conditions. …but cultivated olives maintain high within-species variation and plasticity, enabling adaptation across Mediterranean environments.
- Deciphering the Origins of Commercial Sweetpotato Genotypes Using International Genebank Data. One Brazilian sweetpotato traced back to a CIP accession with a different name, but others did not match anything in the genebank.
- Exploring genetic diversity and selective signatures, a journey through Colombian cassava’s landscape. Colombia’s farmers and environments have shaped its cassava diversity. No word on whether any of it traces back to the CIAT genebank.
- Novel germplasm of tepary and other Phaseolus bean wild relatives from dry areas of southwestern USA. The available genepool for bean breeding gets a welcome boost.
- Insight into root system architecture of buckwheat through genome-wide association mapping-first study. Want drought-resilient, high-yielding buckwheat varieties? Here are the genes — and genotypes — to play with. So the available genepool doesn’t need a boost?
- Non-destructive prediction of nitrogen, iron and zinc content in diverse common bean seeds from a genebank using near-infrared spectroscopy. High-throughput, non-destructive phenotyping methods capture nutritional trait variation across a bean core collection. Wild teparies unavailable for comment.
- Germplasm exploration and digital phenotyping reveal indigenous diversity and farmer preferences in pigeon pea (Cajanus cajan (L.) Millsp.) for climate-smart breeding. Not all phenotyping can be high-throughput, but that doesn’t mean it’s not useful, at least in pigeon peas.
- Agricultural landscape genomics to increase crop resilience. Could have been applied to all of the above, I guess.
Nibbles: Corn diseases, German potato collection, Vietnam rice trials, Endophyte strain, Fish nutrition, Himalayan pea, Subversive seeds
- The US needs better maize.
- German genebank looks for the best potatoes.
- Vietnam looks for better rice in IRRI’s genebank.
- New Zealand markets an endophyte for better grass performance.
- Some Timor-Leste fish are better than others.
- The Himalayas have a better pea. Of some kind.
- How’s that for subversive cataloguing?
A breed is a breed is a breed?
I feel maybe yesterday’s Nibble on the definition of a “breed” may have been a bit too laconic, even for me. So let me give a bit more context.

The link was to a YouTube playlist, which was described thus (link added):
The presentations in this playlist are from the webinar on “Genomic assessment of genetic variation and the future of the breed concept”, originally held on 12.12.2024 under the umbrella of the Food and Agricultural Organization (FAO). This represents the culmination of collaborative work by a diverse group of experts from institutions from all around the world to prepare materials for a sub-chapter in the upcoming 3rd Report on the State of the World’s Animal Genetic Resources for Food and Agriculture.
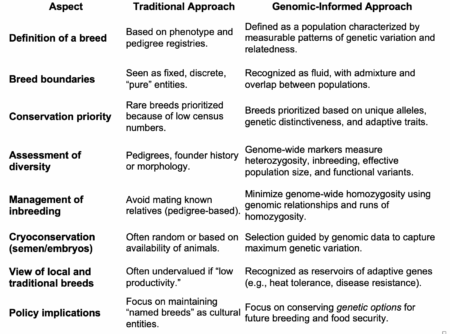
It amounts to over an hour of talks, but if I had to summarize the point the playlist is making, it is that genomics is redefining a breed as a fluid, porous, genotypically-characterized population rather than a fixed, pedigree-based, phenotypic entity. Thus, it is shifting livestock conservation from saving labels (“Breed A”) to preserving genetic options.
Here’s a handy table I came up with to describe the change:
Interesting to juxtapose this to the post a few days ago on how to value and use Indigenous knowledge to solve today’s problems. Would Indigenous livestock keepers necessarily care about those genetic options more than their traditional breed?
It would be great to hear from people engaged in livestock conservation on this. It’s unfortunately not a community I interact with much.
Brainfood: Core collections of…durum, deulkkae, barnyard millet, durian, sesame, flax, Fendler’s horsenettle, jute mallow, barley
- Creation of a core set of durum wheat accessions based on agro-morphological traits with maximum diversity and lower redundancy. From 710 to 13 accessions (2%!) using 32 morphological traits, thanks to Power Core.
- Construction of a core collection of Perilla frutescens (L.) Britton Germplasm in the South Korean gene bank using agro-morphological traits. From 1227 to 235 accessions (19%) using 17 morphological traits, thanks to a bunch of different methods.
- Comprehensive Phenotyping of 1,807 Indian Barnyard Millet (Echinochloa frumentacea Link) Accessions from Indian National Genebank: Unlocking Diversity for Core Set Development. From 1,807 to 271 accessions (15%) using 23 quantitative traits, thanks to Core Hunter 3.
- Genomic resequencing reveals genetic diversity, population structure, and core collection of durian germplasm. From 114 to 26 accessions (23%) using 39 million high-quality SNPs across the genome.
- Development of a composite core collection from 5,856 sesame accessions being conserved in the Indian National Genebank. From 5,856 to 1,768 accessions (30%) using SNPs and phenotypic data.
- Optimizing core collections for genetic studies: a worldwide flax germplasm case study. From 1,593 to 350 accessions (22%) using phenotypic and genotypic data, times 200, thanks to CoreCollection, corehunter III, TrainSel, and more.
- An Optimized Core Sample of the Wild Potato Solanum fendleri in the USA. From 269 accessions, to 38 plants, to 1 accession (0.4%!). Beat that!
- Countrywide Corchorus olitorius L. core collection shows an adaptive potential for future climate in Benin. From 305 to 54 accessions (18%) using 1,114 high-quality SNPs, thanks to ShinyCore. Some indication of usefulness.
- Multi-environmental evaluation of barley core collection against spot blotch for genetic variability and identification of promising genotypes exhibiting resistance. From a core collection of 678 accessions to 2 genotypes that might actually be useful to breeders. Finally!