- The ClimSat classification system—a global climate classification map based on long-term satellite-derived data. There’s a new global climate classification system in town, and it’s better ecologically than Köppen’s.
- The first global agricultural field boundary map at 10 m resolution. Combined with the above, we can now characterize the climate of every agricultural field in the world.
- GEM-Forest: A Global satellite EMbedding–based map of forests and tree crops for 2020. Do any of those fields have tree crops? And how far is the forest?
- Global annual cropland dynamics 2015–2024. The next time we map agricultural field boundaries, there will probably be more of them.
- Climate-induced range shifts support local plant diversity but don’t reduce extinction risk. Those new agricultural fields will be bad for wild plants.
- ‘SiteTool’: a ‘Shiny’ application for field site selection and evaluation. Cool new tool helps you select geographical sites based on ecological characteristics. Could be used to help decide where to collect or evaluate germplasm. Lots of opportunities for combining with some of the above, I suspect.
- Current and future potential of cassava (Manihot esculenta) in Southern Africa: a scoping review. An example of what you can do when you combine different types of spatial (and other data). The area suitable for cassava in Africa will increase, and there’s lots of scope for higher yields too. If we can combine datasets, soon we’ll know which specific fields to grow it in, for higher production, to protect wild biodiversity…
- Global and regional climate modes modulate armed conflict risk. …and to mitigate the risk of conflict.
Nibbles: Pearl millet redux, Garden plants, Armenian pics, Seeds galore, Heavenly Book, Pastoralism threats
- Pearl millet is getting the hybrid treatment. And, loving it.
- Want to know what to grow in your garden? Yes, even pearl millet.
- Nice pics of Armenian landscapes, food and foodways. No pearl millet in sight.
- The latest monthly newsletter from The Botanist in the Kitchen does seeds. Pearl millet unavailable for comment.
- China is genotyping and phenotyping (almost) everything. Pearl millet feeling left out.
- If pearl millet fails, there is always pastoralism. No, wait…
Brainfood: Targets, Plant Treaty, Decolonization, Fonio germination, Recalcitrant seeds, Microbiome, Taro seed system
- Status and future of seed conservation of threatened plants in the post-2020 era. 21% of threatened plants are conserved in genebanks across 44 countries in Europe and western Asia. Not bad, but not good enough. I wonder how many of those 21% will be of interest to breeders?
- How the international treaty on plant genetic resources for food and agriculture can support effective germplasm exchange: four Colombian case studies. The Plant Treaty can really help a country’s genebanks and breeders drive agricultural development, given half a chance.
- Reconciliation or re-colonization? Critical perspectives on seed banking and colonialism. Indigenous communities need to be careful in collaborating with genebanks and breeders.
- Impacts of climate change on fonio millet: seed germination ecology and suitability modelling of an indigenous West African cereal. Climate change will screw up the germination of fonio in some places, so genebanks and breeders better get cracking.
- Euterpe edulis seed recalcitrance: difficult, yes, but not impossible to genebank. Tricky seed storage behaviour need not deter genebankers.
- Accelerated aging caused diversity and specificity loss in the bacterial communities of Brassica napus seedlings. Genebanks should be careful with their seed aging experiments, because they might screw up the seed microbiome.
- Understanding Biotic Constraints to Taro (Colocasia esculenta) Production in the Derived Savanna and Humid Forest Agroecosystems of Nigeria. Genebanks need seed systems though.
Gaps galore in collards collections
Quick follow-up to my post a few days ago on the recent study of the origin of the collard greens grown in the Moroccan oases of the Draa and Ziz valleys.
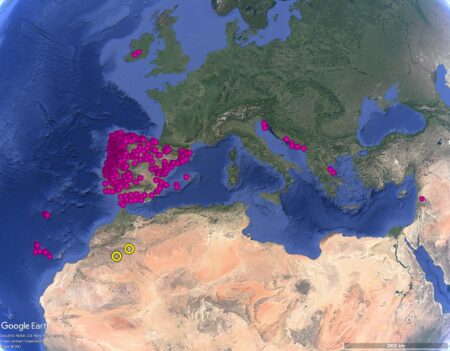
Ethnobotanists Bronwen Powell and Abderrahim Ouarghidi used historical texts, linguistics, and Indigenous knowledge in their investigation, but of course it’s also possible to use genetics to figure out where the plants may have came from. Especially as there are plenty of accessions labelled Brassica oleracea var. acephala in the genebanks that share their data on Genesys — just over 1500 in fact.
Alas, that might in practice turn out to be tricky, though, due to the somewhat — ahem — skewed geographic distribution of the accessions in question. The yellow circles in the map below show the approximate locations of those oases on the edge of the Sahara.
Still worth trying, in my view, but really more than anything this should be an encouragement to do some more collecting. Or get more genebanks on Genesys. Or identify more B. oleracea accessions to variety level. Or…
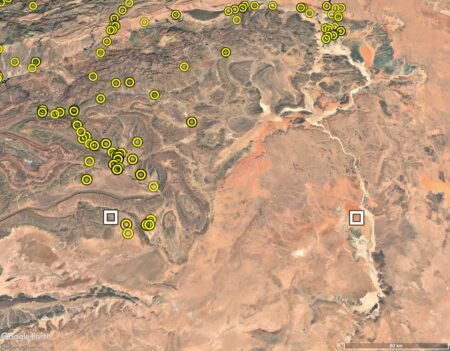
What else has been collected in the Draa and Ziz valleys or thereabouts? Surprisingly little, mainly wheat, barley, chickpea, faba bean and alfalfa. The general location of the valleys is now shown by white squares.
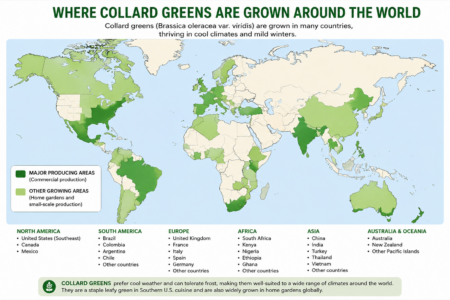
Incidentally, the. map below is where ChatGPT thinks collards are grown around he world. I really have no idea how accurate it is. I hope someone will tell us.
Nibbles: Crop mapping, Climate change impacts, Rice cheese, Andean blueberry, Rare apples, Hungarian genebank, Old seed collection
- AI doesn’t recognize tropical agriculture very well.
- So presumably it can’t easily be used in assessing climate change impacts in agricultural heritage systems? FAO has some ideas on how to do it.
- Maybe rice heritage systems can be used to make cheese.
- I bet Andean blueberry (Vaccinium floribundum) goes great with rice cheese.
- But if not, heritage apples will probably do.
- The Hungarian genebank is hoping to inject heritage grains into non-heritage agricultural systems. AI and FAO unavailable for comment.
- Maybe AI can help with the mystery of this old seed collection at the Natural History Museum, London.