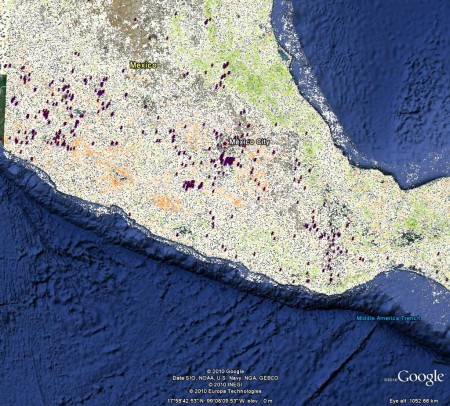
I’ve been exploring Google’s 3d trees thing a bit, to work out just how cool it is. I said in my previous post on this that it could eventually be used to document and virtually explore field genebanks (of coconuts, say, or breadfruit). But of course you can explore a few forests around the world right now, so I wondered if any crop wild relatives have been collected in any of these places.
I’ve been exploring Google’s 3d trees thing a bit, to work out just how cool it is. I said in my previous post on this that it could eventually be used to document and virtually explore field genebanks (of coconuts, say, or breadfruit). But of course you can explore a few forests around the world right now, so I wondered if any crop wild relatives have been collected in any of these places.
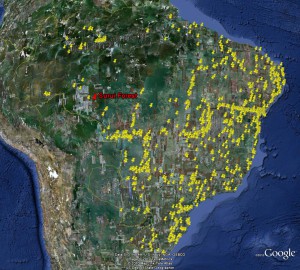 The answer is, alas, no, at least for Surui Forest in Brazil, one of the couple of “wild” places for which Google currently has 3d trees (the others are urban areas). At left you can see the distribution of accessions of crop wild relatives in Brazil, according to Genesys. Unfortunately, none fall within the area for which Google has 3d data.
The answer is, alas, no, at least for Surui Forest in Brazil, one of the couple of “wild” places for which Google currently has 3d trees (the others are urban areas). At left you can see the distribution of accessions of crop wild relatives in Brazil, according to Genesys. Unfortunately, none fall within the area for which Google has 3d data.
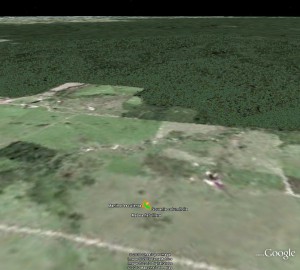 I did get a hit in GBIF for a cultivated cassava just outside the forest. But that’s not quite the same, I agree. Oh well, maybe we’ll soon have more data. 1
I did get a hit in GBIF for a cultivated cassava just outside the forest. But that’s not quite the same, I agree. Oh well, maybe we’ll soon have more data. 1