- Wild camel genetically distinct from the domesticated kind. Well I never.
- Maya tapped into their “sacred groves” to build temples, which did not end well.
- Boffins extract DNA from ancient barley in Upper Egypt, find it was 2-rowed, but derived from a 6-rowed ancestor. No word on whether it was used to make beer, but my guess is yes.
- Large Y chromosome microsatellite study of Eurasian cattle does “not support the recent hypothesis on the origin of Y1 from the local European hybridization of cattle with male aurochsen.” This could run and run.
- I like this idea: a garden of poisons.
- Agroforestry’s coming-of-age party coming up. You going? Let us know.
- Multiple explanations for lactase persistence.
Ex situ redux
After a period in which ex situ conservation has been downplayed by the conservation community (except for agrobiodiversity where it is still the main conservation strategy) ex situ conservation is now widely accepted as an increasingly necessary complement to in situ forms of conservation (IUCN 2002; BGCI 2000), especially protected areas (e.g. Abanades García & al. 2007).
That’s from a new report for the Council of Europe entitled “The impacts of climate change on plant species in Europe,” prepared by Prof. Vernon Heywood of the School of Biological Sciences, University of Reading, with contributions by Dr Alastair Culham. You’ll find it on p. 39 after a very thorough review of the issues. Nice to see such a bold statement. The report is one of several prepared for the Group of Experts on Biodiversity and Climate Change of the Convention on the Conservation of European Wildlife and Natural Habitats. Thanks to Danny for the tip.
Nibbles: Cheese, Dog genetics, Olives on Crete, Polyploidy, Pollination
- Making French cheese in the Himalayas.
- The latest on how to build your perfect dog.
- “The scientists are putting the all the trees which must be saved into a data bank.” Clever scientists.
- Polyploidization so, so much more than merely the sum of genomes.
- “The expected direct reduction in total agricultural production in the absence of animal pollination ranged from 3 to 8%…” Thank goodness for Operation Pollinator, eh?
Nibbles: Dahlias, Perennials
- Dahlias: good to look at, good to eat.
- Why agriculture bypassed herbaceous perennials, until now.
Monitoring plants of “Community interest” in Europe
There’s been an item in the news the last couple of days to the effect that “[a] report by the European Commission shows that habitat and wildlife protection targets across Europe will be missed…” Digging a bit deeper into that seemingly simple statement led me to a hitherto unknown (to me) world of EU rules and regulations and reporting requirements.
Let’s start at the beginning. There’s a thing called the Habitats Directive (1992). This requests all Member States “to monitor habitat types and species considered to be of Community interest.” It’s unclear to me how they were selected (perhaps someone out there can tell us), but these species are listed in various annexes to the Directive, though that sounds more simple than it is:
Where a species appears in this Annex but does not appear in either Annex IV or Annex V, the species name is followed by the symbol (o); where a species which appears in this Annex also appears in Annex V but does not appear in Annex IV, its name is followed by the symbol (V).
Anyway, Article 17 provides for regular reports on implementation of the Directive, and the report “for the period 2001-2006 for the first time includes assessments on the conservation status of the habitat types and species of Community interest.”
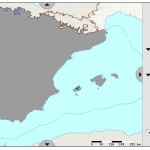
The website which houses the Article 17 reports is, well, complicated, but well worth exploring. The most interesting bit from an agrobiodiversity perspective is the page from which you can get species reports. These include all kinds of information about the status of those “species considered to be of Community interest,” country by country (there’s also an overall summary). Some of these species “of interest” are crop wild relatives such as Allium grosii, an endemic to the Balearic Islands (click the map to enlarge it).
There’s a few more CWRs in those annexes, though not all that many. A Hungarian Pyrus, for example. Any chance to get a few more on there? The bureaucratic infrastructure and mechanism for regular monitoring and early(ish) warning of any threats would seem to be well and truly in place, European Union-style.