- Potato and Food Security in China. Huge expansion, mainly due to product diversification, but still room for growth. But how will it end? Like bananas?
- Converging phenomics and genomics to study natural variation in plant photosynthetic efficiency. Chlorophyll fluorescence technologies are revolutionizing phenotyping. Now everyone will want another gadget.
- Is DNA fingerprinting the gold standard for estimation of adoption and impacts of improved lentil varieties? It’s not about yield.
- A florigen paralog is required for short-day vernalization in a pooid grass. Nope, I can’t say it better than the press release: Ancient gene duplication gave grasses multiple ways to wait out winter.
- Drones for Conservation in Protected Areas: Present and Future. Sure, why not. On-farm too?
- Genome-Enhanced Detection and Identification (GEDI) of plant pathogens. Sort of barcoding for bugs.
- Self-domestication in Homo sapiens: Insights from comparative genomics. There’s a domestication syndrome for humans too.
- Cryopreservation of Citrus limon (L.) Burm. F Shoot Tips Using a Droplet-vitrification Method. Well, at least two varieties work.
- Farmers Drive Genetic Diversity of Thai Purple Rice (Oryza sativa L.) Landraces. Well, who else?
- Genetic Diversity of Ethiopian Tef [(Eragrostis tef (Zucc.) Trotter] Released and Selected Farmers’ Varieties along with Two Wild Relatives as Revealed by Microsatellite Markers. The landraces are distinct from the released varieties, and more diverse.
- Biodiversity Observations Miner: A web application to unlock primary biodiversity data from published literature. Nice enough, but you need to upload a PDF corpus. Why not let it loose on the internet?
- Cross-species hybridization and the origin of North African date palms. I always knew that P. theophrasti would come in useful.
- Revisiting the versatile buckwheat: reinvigorating genetic gains through integrated breeding and genomics approach. Start with a database, core collection, and wild relatives. Gratifyingly old-fashioned.
- The genome of broomcorn millet. That would be Panicum miliaceum.
Nibbles: A2S2019, ICRISAT seeds, High protein rice, American grapes, Religion & diet, Australian NUS
- Access to Seeds comes out with 2019 edition. Not much change, alas. LATER: Ok, I stand corrected. And spanked.
- Maybe this ICRISAT online seed tool will help with that access.
- How many people will have access to this high protein rice?
- Indigenous American grape species: more than just rootstocks.
- The religion of diets.
- Bush tucker is a religion for some.
Nibbles: Heirlooms double, Seed huntress, Sequencing, ABS
Detecting Wisconsin’s Wild Cranberries from Space
This is a guest post by Vanesa Martin, Anastasia Kunz, Nicole Pepper and Eli Simonson of the NASA DEVELOP Colorado office. Many thanks to all of them. And thanks also to Colin Khoury for helping to make it happen.
NASA DEVELOP, as a part of NASA’s Applied Sciences Program, addresses environmental issues through interdisciplinary research projects that apply the lens of NASA Earth Observations to community concerns around the globe. Over the last year, researchers at the USDA ARS National Plant Germplasm System (NPGS) in Fort Collins, Colorado approached the NASA DEVELOP program to determine whether NASA satellites could be used to monitor and map Crop Wild Relatives (CWR) more efficiently than standard field methods alone. This collaboration has created a push to experiment with the mapping of different types of CWRs, in the hope of eventually producing an operational method to accurately monitor CWRs globally.
In the fall of 2018, our NASA DEVELOP team was formed to focus efforts on using satellite information to map the distributions of wild cranberries within a Landsat scene in northern Wisconsin. Previously, the USDA ARS’s method for mapping species distribution had exclusively considered bioclimatic and environmental factors, and these provided only a rough prediction of possible cranberry presence that looked like this:
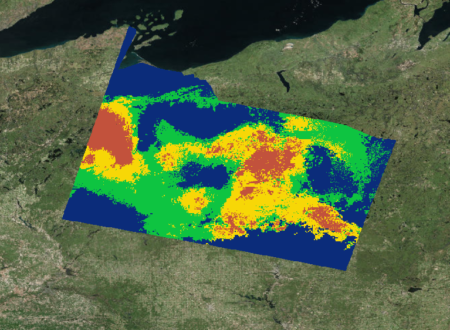
With the aim of narrowing down the areas of predicted presence, the team used cranberry presence data from publicly available databases like GBIF and BISON to train initial habitat distribution models through Software for Assisted Habitat Modeling (SAHM). In addition to publicly available cranberry presence data, we incorporated bioclimatic and topographic variables derived from WorldClim (a global climate database with 1 km resolution) to mirror the current practices of USDA. We later incorporated ClimateNA data (a national climate database with 30m resolution) to assess whether our distribution maps improved.
The publicly available presence data we used was not suitable for remote sensing purposes, given that remote sensing requires highly accurate location information where the spectral signatures of the target species is clearly discernible. Consequently, we generated our own predicted presence points in order to incorporate spectral data and NASA Earth Observations into these models. These user-generated points, which we based on research we did of our two target cranberry species, were sent to our field cranberry experts to verify that they actually represented sites of probable cranberry presence.
After getting their approval and employing the user-generated points, we incorporated spectral detection into the models, creating maps based on the detection of the spectral signature of these species instead of predicting their possible locations based on suitable environmental conditions. This was the result, with red being the likeliest locations of cranberry presence:
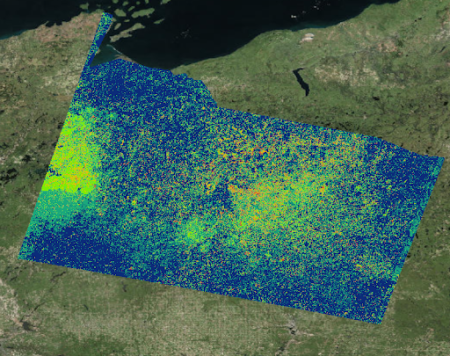
One way we assessed the accuracy of our detection method was by overlaying a commercial cranberry layer we obtained from the Wisconsin Department of Natural Resources on to our maps to see how well they aligned with our binary detection maps. The binary detection maps did indeed locate commercial cranberry crops, despite the fact that we did not use any commercial cranberry presence points to train our models. With this assurance in hand, we felt more confident in sending our final incorporated maps to our field partners and experts for a final verification.
We hope to be able to continue working with our partners to conduct in-field verification of our maps. For now, however, it’s apparent that spectral data can positively influence the research efforts of our partners at the USDA ARS, and their larger goals of improved food security and biodiversity.
Brainfood: Intensification, Yemen ag, Czech barley, Bangladesh community genebank, Agrobiodiversity Index, North American CWR, Israeli genebanks, Biofortified wheat, QDS, Collecting Miscanthus, Ethnobotany, NUS, Pecan diversity, Korean ponds, CWR gaps double, Salty rice
- Agricultural intensification, dietary diversity, and markets in the global food security narrative. Intensification is all well and good but it needs to be sustainable and nutrition-sensitive.
- Health, Seeds, Diversity and Terraces. Maybe evolutionary plant breeding can help with that.
- Identification of barley powdery mildew resistances in gene bank accessions and the use of gene diversity for verifying seed purity and authenticity. It’s difficult to deal with heterogeneous accessions.
- The USD 1,875.95 Seed Center. A serious-looking community seed bank in Bangladesh.
- Assessing agroecosystem sustainability in Cuba: A new agrobiodiversity index. Not same as the old index.
- North American Crop Wild Relatives, Volume 1. Volume 1?
- Ex-situ conservation strategies for endangered plants in the Israel Gene Bank. Not just crops, and not just conservation…
- The Institute of Evolution Wild Cereal Gene Bank at the University of Haifa. …and not even the only genebank in Israel.
- Assessing Genetic Diversity to Breed Competitive Biofortified Wheat With Enhanced Grain Zn and Fe Concentrations. Four translocations from rye and various Aegilops species have resulted in 8 biofortified bread wheat varieties after a decade of work. Compare and contrast with potatoes.
- Improving efficiency of seed system by appropriating farmer’s rights in India through adoption and implementation of policy of quality declared seed schemes in parallel. FAO’s Quality Declared Seed (QDS) system is the way to go.
- Collecting wild Miscanthus germplasm in Asia for crop improvement and conservation in Europe whilst adhering to the guidelines of the United Nations’ Convention on Biological Diversity. It can be done.
- Making friends in the field: How to become an ethnobotanist – A personal reflection. Yes, it can.
- Mainstreaming Underutilized Indigenous and Traditional Crops into Food Systems: A South African Perspective. Start by having researchers translate their findings for policy makers.
- Genotyping by sequencing (GBS) and SNP marker analysis of diverse accessions of pecan (Carya illinoinensis). Geographic patterning of genetic diversity and SNPs for dichogamy found.
- Trait-based evaluation of plant assemblages in traditional farm ponds in Korea: Ecological and management implications. Dumbeongs are carefully managed. Well there’s a shocker.
- Conservation gap analysis of crop wild relatives in Turkey. There still are some.
- An in situ approach to the conservation of temperate cereal crop wild relatives in the Mediterranean Basin and Asian centre of diversity. 10 locations would do.
- Molecular characterization and identification of new sources of tolerance to submergence and salinity from rice landraces of coastal India. 5 of 98 accessions had novel alleles.