Roderic Page has a perceptive, amusing rant over at iPhylo today on why people who come up with phylogenetic trees or cluster diagrams, say as a result of a fancy molecular study, don’t routinely archive them in TreeBASE. His answer is threefold:
1. It is not at all obvious that databasing trees is useful
2. The databases we have suck
3. There’s no obvious incentive for the people producing trees to database them
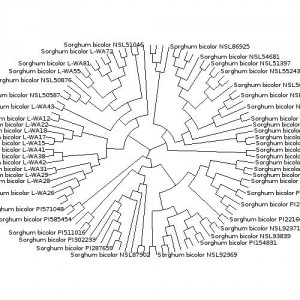 Having spent an hour or so with TreeBASE trying to get the diagram for cultivated sorghum reproduced here, I can certainly sympathize with point number 2. I can’t say much about “the underlying data model, the choice of programming language, the use of a Java applet to display trees” or “the voluminous XML output”, but “the Byzantine search interface” certainly contributes to TreeBASE being “a bag of hurt.”
Having spent an hour or so with TreeBASE trying to get the diagram for cultivated sorghum reproduced here, I can certainly sympathize with point number 2. I can’t say much about “the underlying data model, the choice of programming language, the use of a Java applet to display trees” or “the voluminous XML output”, but “the Byzantine search interface” certainly contributes to TreeBASE being “a bag of hurt.”
And yet I’m not so sure about Dr Page’s point number 1. I have a feeling that a way of storing and comparing diagrams illustrating the genetic relationships among genebank accessions or the wild relatives of a crop (including genepool concepts) might well be welcome in the agrobiodiversity community. Which would render point 3 moot. At least if it wasn’t TreeBASE. But don’t let me speak on your behalf. If you have a strong opinion, one way or another, leave us a comment.
LATER: If not TreeBASE, then perhaps OneZoom?
One Reply to “Do we need an archive of cluster diagrams?”