The main reason for my quick trip to Papua New Guinea last week was to get up to date on Bogia Coconut Syndrome (BCS). Readers with a long(ish) memory may remember that we blogged about this some time back. Quick recap. BCS is a phytoplasma disease first reported about 20 years ago in Yaro Plantation, near Bogia, Madang Province, on the northern coast on PNG.
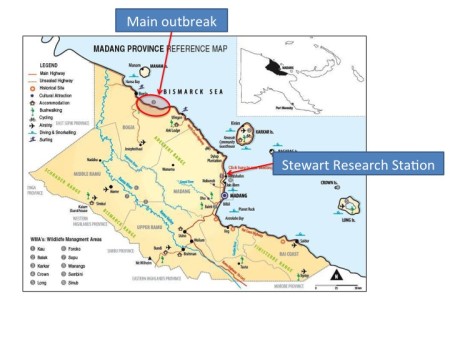
Since then, it has devastated coconuts in a large contiguous area SE of Bogia, but it has also leap-frogged to a number of sites further along the coast towards Madang and beyond. In 2013, it was spotted in Mobdub, which is only a few kilometres from the Stewart Research Station of PNG’s Cocoa & Coconut Research Institute (CCI), home of COGENT’s International Coconut Genebank for the South Pacific (a collection placed under Article 15 of the International Treaty through a tripartite agreement involving the PNG government, Bioversity on behalf of COGENT and FAO). Here’s Alfred Kembu, the curator of the collection, talking about it in a video by Roland Bourdeix, who used to be the coordinator of COGENT, the global coconut network:
https://www.youtube.com/watch?v=wSADJB4tKXEThere are more videos on the disease and the threat it poses to the collection by COGENT and CIRAD. The closest outbreak to the collection is now about 2 km away. There has been a ban in place in Madang for 3 years now on the movement of nuts that have not been de-husked or are sprouting, as the pattern of movement suggests human agency.
At one of these outbreak sites 1, first spotted in 2009, the affected area went from a radius of 200 m in 2011 to 1800 m in 2013; all palms are expected to be dead there within 2-5 years. The disease can kill 1-5 palms per month, meaning that a 1 ha plantation can be destroyed in 4 years. This is an extremely dangerous disease for local smallholders, as well as for the genebank. The same phytoplasma is suspected to affect other palm crops, such as oil palm and betel nut, but also banana and possibly others. This photo shows palms at different stages of development of the disease. In the final stage, nothing but the stem is left.
I went to Madang to take part in the inception workshop of the ACIAR-funded project “Bogia Coconut Syndrome in Papua New Guinea and related phytoplasma syndromes: Developing biological knowledge and a risk management strategy,” which is led by Prof. Geoff Gurr of Charles Sturt University, Queensland, Australia. The project activities, which will last four years, center on intensive and repeated surveying and sampling of both plants and insects in a number outbreak areas (in particular for DNA analysis using “loop-mediated isothermal amplification,” or LAMP, which is apparently a method of identifying phytoplasma DNA which is more efficient and cheaper than standard PCR), followed by a series of transmission experiments.
Although the main output will be basic scientific information on BCS (its causal organism, location in the host plant(s), possible alternate hosts, vector(s) etc, none of these are currently entirely clear), a management plan for the disease will also be devised on the basis of the results. The two-day workshop focused on discussing methodological and logistical issues pertaining to the project, which will involve a number of other PNG institutes apart from CCI, but we also talked about the possible relocation of the germplasm collection to a safer locality. Not quite ready for that yet, but we’ll keep working on it. Here is a shot of part of the collection, by the way: talls on the left, dwarfs on the right, cacao in the understory, which is sold to generate some income to help keep the collection going.
We’ll keep you posted…


“The same phytoplasma is suspected to affect other palm crops, such as oil palm and betel nut” this statement by Luigi to suspect BCS in oil palm is not true. This should read “sago palms and confirmed in betelnuts”.